Automated 3D / 4D / 5D segmentation
Analyze microglia, neurons, macrophages, immune cells, and other motile cells across single-channel or dual-channel time-lapse microscopy datasets.
neuroGlia5D is built for microglia, neurons, macrophages, and other dynamic cell populations. It combines automated segmentation, object-centric review, and interactive correction without forcing you into an HPC workflow.
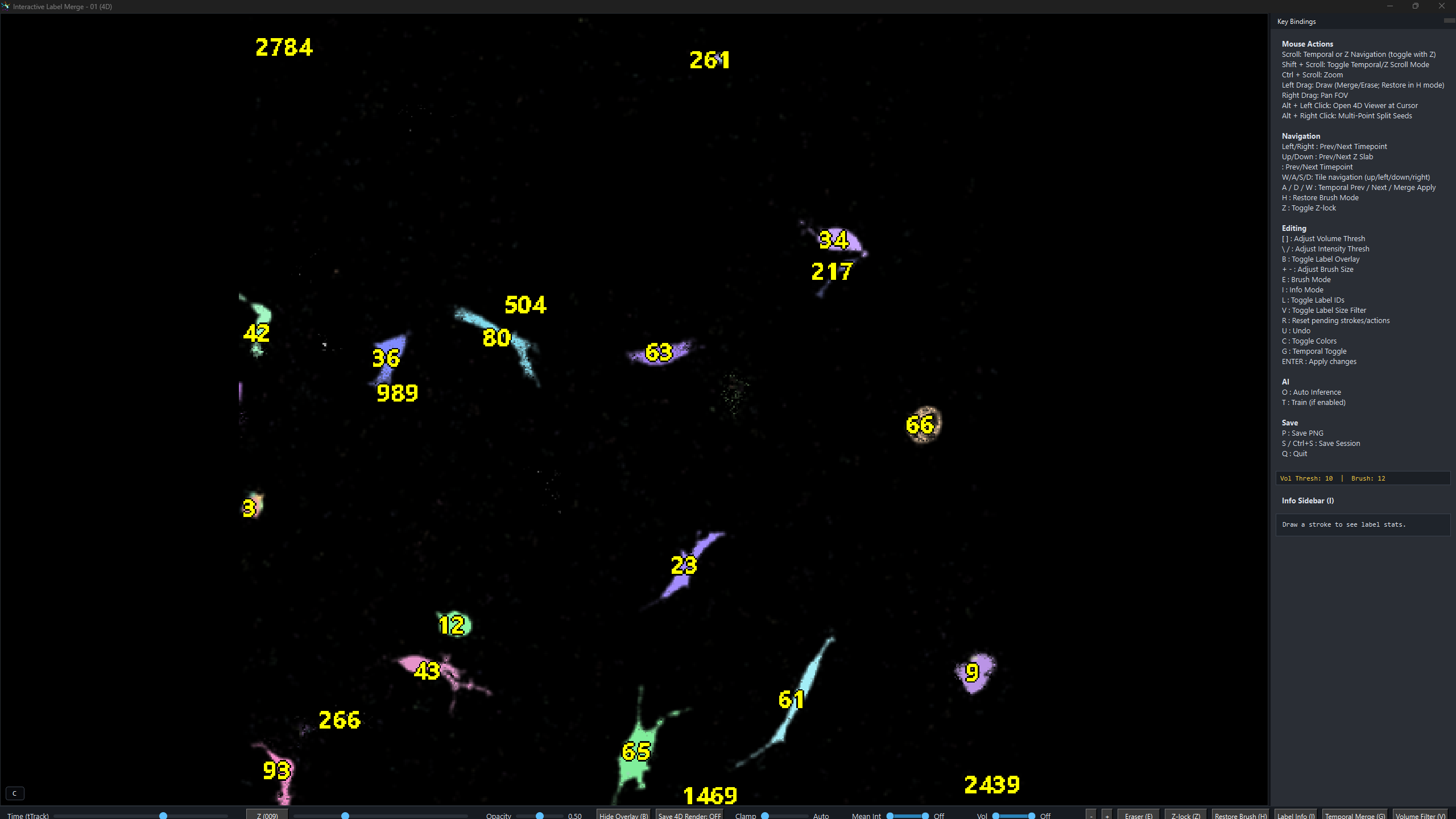
Analyze microglia, neurons, macrophages, immune cells, and other motile cells across single-channel or dual-channel time-lapse microscopy datasets.
Navigate persistent labels across time, inspect cell identities, and correct local segmentation issues without exporting to a separate manual curation stack.
Designed around GPU memory behavior, adaptive tiling, and streaming so large datasets remain usable on an NVIDIA-equipped Windows laptop or workstation.
Live microscopy often forces a painful tradeoff between field of view, temporal resolution, and downstream usability. Existing workflows frequently zoom into small regions just to preserve dynamic detail, then lose the broader biological context that actually matters.
neuroGlia5D was built to reduce that compromise. It treats segmentation, navigation, and rendering as one connected system so users can move from whole-volume context to individual cellular structures without rebuilding the workflow around a single tiny ROI.

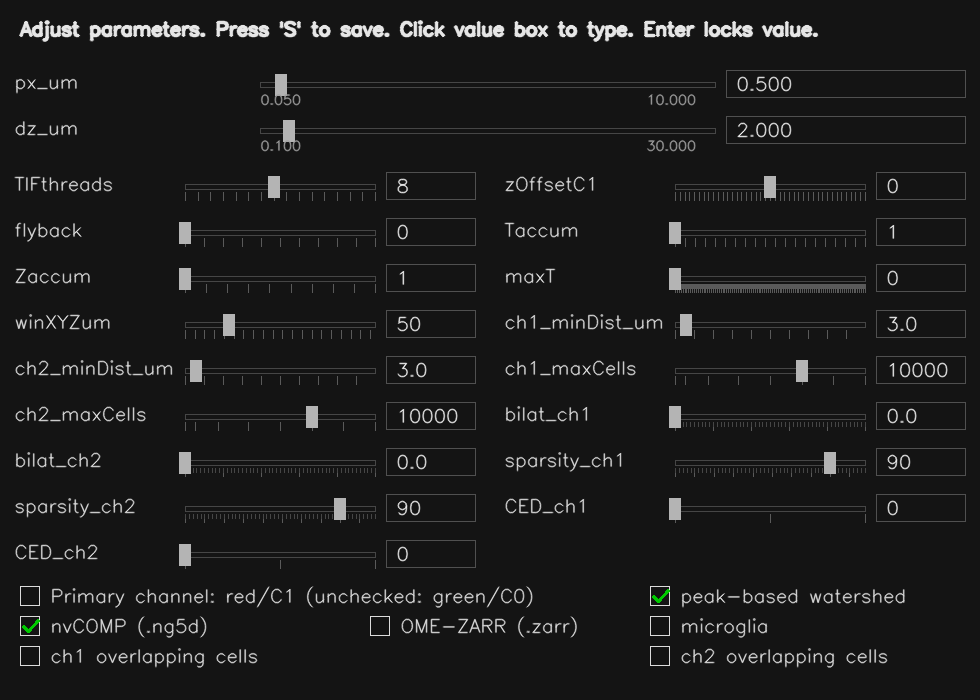
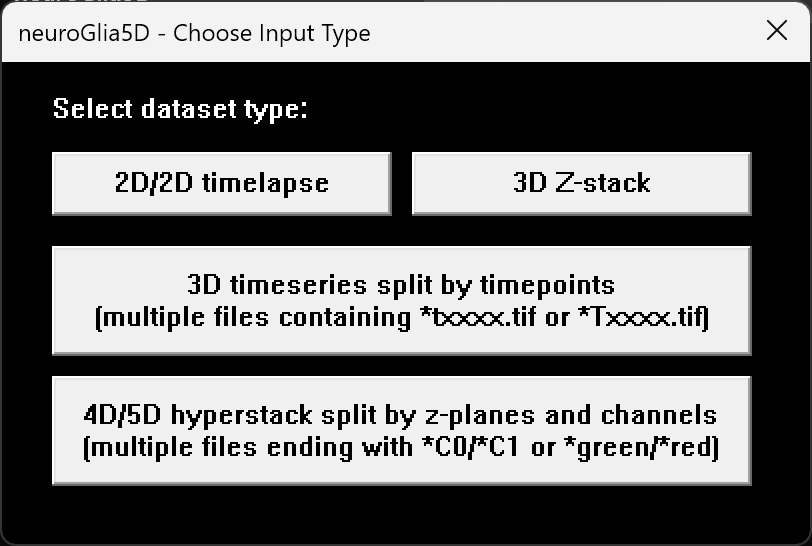
Use the workflow page for step-by-step screenshots, or review pricing first if you need procurement-ready information before starting a trial.
neuroGlia5D is built around GPU-native algorithms rather than a loose collection of CPU-era tools. Data access, filtering, resampling, segmentation, and topology tracking were designed together with GPU execution and memory constraints in mind.
Instead of treating labels as a disposable overlay, the system models each segmented cell as a persistent 4D object. That makes navigation, review, interaction analysis, and downstream AI-assisted correction more coherent than frame-by-frame workflows.
The comparison below describes product focus rather than absolute rankings. Cell5D is not affiliated with, endorsed by, or sponsored by the third-party products named here.
GPU-native software for 3D, 4D, and 5D segmentation and analysis of live biological cells from fluorescence microscopy.
No. It is designed for a Windows laptop or workstation with a modern NVIDIA GPU.
Multi-page TIFF, OME-TIFF, and OME-ZARR input, with output options including .ng5d, .ome.zarr, and .tif.
It was built around microglia-neuron workflows and is also aimed at macrophages, immune cells, neurons, and other dynamic cell populations.
Yes. Start with the trial, the workflow guide, or the pricing page for internal review and purchasing.
Yes. The site already states that universities can be invoiced via purchase order.
Email cell5d@outlook.com for dataset questions, live demos, procurement support, or performance discussion. Tutorials and product examples are on YouTube @neuroGlia5D.